Understanding biology is no longer just about knowing which proteins exist in a cell — it’s about knowing where they function.
Proteins do not act in isolation. They assemble, interact, and signal within highly organized subcellular neighbourhoods. When these spatial dynamics are disrupted, disease often follows. Many neurological disorders, cancers, and inflammatory conditions are linked not only to protein dysfunction, but to abnormal protein localization.
Yet, despite decades of progress in proteomics, mapping proteins to precise cellular locations has remained a significant challenge.
Why Location Matters in Proteomics
Traditional proteomics excels at depth and coverage, but it typically averages signals across whole cells or tissues. This approach makes it difficult to understand region-specific biology, such as:
- Protein composition at synapses
- Signaling hubs near cell membranes
- Molecular interfaces between organelles
- Nanoscale structures like Lewy bodies or receptor clusters
On the other hand, imaging-based techniques reveal cellular architecture but often sacrifice unbiased discovery, molecular breadth, or nanoscale resolution. To truly understand cellular function, molecular identity must be connected to spatial context.
Bridging Imaging and Proteomics
Recent advances in microscopy-guided proteomics are beginning to close this gap.
By combining high-resolution imaging with controlled chemical labeling, researchers can now isolate and analyze proteins from specific regions of interest (ROIs) within cells or tissues. This approach enables scientists to study localized proteomes rather than bulk averages — unlocking insights that were previously inaccessible.
Moving Beyond Optical Limits
While optical microscopy defines where researchers look, chemistry determines what gets labeled.
For many subcellular structures, optical resolution alone is sufficient. However, when studying nanoscale architectures — such as synaptic nanoclusters, mitochondrial contact sites, or receptor interactomes — additional precision is required.
Advances in photochemical labeling strategies now allow protein tagging to be confined not only by microscopy, but also by controlled reaction distance, enabling molecular resolution at the nanometer scale.
This dual confinement — spatial selection by imaging and molecular selectivity by chemistry — represents a major shift in spatial proteomics.
Spatial Precision Across Biological Systems
Microscopy-guided proteomics is transforming research across multiple contexts:
Cellular Models
Researchers can now:
- Quantify localized signaling events
- Map drug-responsive microdomains
- Study organelle- or sub-organelle–specific proteomes Complex Tissues In structured environments such as tumors, brain tissue, or organoids, spatial proteomics reveals:
- Cell–cell interaction zones
- Signaling gradients within tissue architecture
- Disease-driven remodeling at biological interfaces This added spatial layer provides a deeper understanding of how biological systems are organized and how they change in pathology.
From Mechanism to Medicine
For translational and pharmaceutical research, spatial precision is critical. Many therapeutic targets operate within highly localized molecular domains, not across entire cells. Identifying druggable vulnerabilities, off-target interactions, or spatial biomarkers requires tools capable of resolving biology at the scale where function occurs. Microscopy-guided proteomics enables scientists to:
- Define the proteome of a specific cellular feature
- Track how it changes in response to treatment
- Generate hypotheses for new targets and biomarkers
This transforms exploratory studies into structured, discovery-driven pipelines.
Two Complementary Strategies for Discovery
Modern spatial proteomics benefits from a two-tiered approach:
Broad Discovery
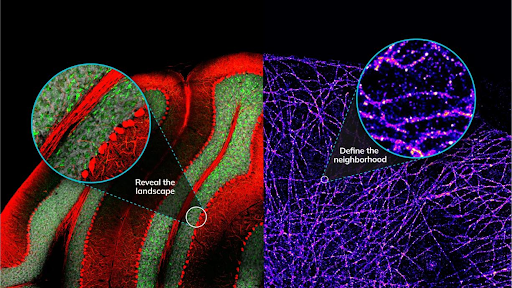
An initial, unbiased exploration captures the full proteomic landscape of a defined region, revealing which proteins are present and how they respond to perturbation.
Focused Precision
Targeted strategies then zoom in on specific proteins or complexes, isolating their immediate molecular neighborhood at nanoscopic resolution.
Together, these approaches allow researchers to move seamlessly from landscape-level discovery to mechanistic insight, using a single integrated workflow.
Redefining Proteomic Insight at the Nanoscale
As biology increasingly focuses on spatial organization, nanoscale proteomics is becoming essential.
The ability to define proteomes at tens-of-nanometers resolution reshapes how scientists investigate:
- Synaptic signaling
- Cancer cell communication
- Immune–tumor interfaces
- Drug mechanism of action
By uniting imaging, chemistry, and mass spectrometry, spatial proteomics reveals not justwhat proteins are present — but where biology truly happens.
A New Paradigm for Discovery
Spatial proteomics is not simply an incremental improvement. It is a paradigm shift.
It enables researchers to ask sharper questions:
- What defines this functional niche?
- Which proteins assemble or disperse here under disease or treatment?
- How do therapies rewire nanoscale networks?
At the intersection of cell biology and proteomics, nanoscale discovery is redefining how we uncover mechanisms, biomarkers, and therapeutic opportunities.


test