Sequencing in Epigenetics: ChIP-seq
This month, we will be covering an executive overview in one of the key sequencing methods in epigenetics, ChIP-seq. This method is employed by researchers to further understand and map protein-DNA interaction of transcription factors onto the genome, providing another layer of data into multi-omics research to further resolve biological mechanisms and enabling disease marker discoveries.
The Evolution of ChIP-seq: From Foundational Mapping to Integrated Discovery
Understanding the trajectory of key applications is crucial for assessing market needs, ROI, and future-proofing your NGS investment. Chromatin Immunoprecipitation followed by sequencing (ChIP-seq) is a paradigm case—a technique whose applications have defined entire fields and continue to evolve.
What is ChIP-seq?
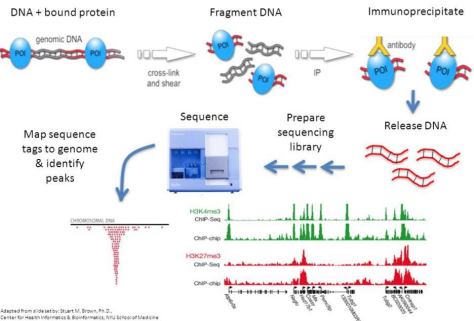
ChIP-seq is a method for sequencing DNA that are bound by proteins. The method involves:
1. Cross-linking bound protein of interest (POI) with its DNA target
2. Genomic DNA isolation and fragmentation
3. Immunoprecipitation via antibody binding with POI
4. Purify antibody-captured POI with magnetic beads
5. Remove DNA-POI cross links
6. Library prep
7. Sequencing
8. Analysis by mapping read counts to genome. Read counts that peak in a region indicate POI binding activity
Part 1: Historical Applications – Laying the Genome Regulation Blueprint
Since its emergence in the mid-2000s, ChIP-seq revolutionized molecular biology by providing the foundation for high-resolution, genome-wide map of protein-DNA interactions. Its core historical applications established the understanding of the epigenetic and regulatory code:
1. Transcription Factor (TF) Binding Site Discovery: Replacing low-throughput methods like ChIP-chip (microarray based), ChIP-seq became the gold standard for identifying where TFs bind, enabling the systematic reverse-engineering of gene regulatory networks controlling development, cell identity, and response to stimuli.
2. Histone Modification Mapping: By mapping modifications like H3K4me3 and H3K27ac, researchers could identify “tightly” or “loosely” packed chromatin regions, implicating transcriptional accessibility and potentially annotate functional genomic elements across the genome, moving beyond the coding sequence to understand the regulatory mechanisms.
3. Epigenetic Profiling in Development & Disease: ChIP-seq allowed comparative studies between cell states, revealing how dynamic epigenetic
landscapes guide differentiation and how their dysregulation contributes to diseases like cancer (e.g., identifying oncogenic enhancers or repressive domains).
4. RNA Polymerase Mapping: It enabled the study of transcription initiation and elongation by pinpointing the location and status of Pol II, linking regulatory events directly to transcriptional output.
5. Large-Scale Consortium Science: ChIP-seq was the methodological backbone of atlas-scale projects like ENCODE and Roadmap Epigenomics Project, which aimed to create comprehensive reference maps of regulatory elements for the human and model organism genomes.
In this era, the primary value proposition for NGS was throughput and resolution— enabling these genome-wide discoveries that were previously impossible.
Transition: The Maturation and the Need for Integration
While the historical applications remain vital, the ChIP-seq field matured. The question shifted from “Where does this protein bind?” to “How does this binding event function within a complex, multi-layered regulatory system to produce a phenotypic outcome?”
Standalone ChIP-seq data provided a static, one-dimensional protein binding regions map, limiting its mechanistic and translational power. This created a clear market pull for NGS platforms and solutions that could support multi-omics integration and higher resolution analyses within complex tissues. The technique itself began to evolve towards new methods addressing its key limitations: high cell input requirements, noisy backgrounds, and the inability to capture cellular heterogeneity.
Part 2: Current-State & Latest Application Trends – The Integrated, Functional, and Single-Cell Era
Today, ChIP-seq’s greatest value is realized not in isolation, but as a core component in a convergent analytical pipeline. The latest trends address the needs of translational research and drug discovery, creating specific demands on NGS capabilities.
Current State – Integrated Omics:
The state-of-the-art application is correlative and integrative analysis. ChIP-seq data is now layered with other data such as:
• ATAC-seq (for open chromatin) to distinguish functional binding. • RNA-seq to link regulatory changes to gene expression.
• Hi-C-seq to connect linear binding events with 3D chromatin looping. • WGBS to overlay DNA methylation patterns.
This integration is essential for biomarker discovery and target validation key medical fields such as oncology, neurology, and immunology. For NGS stakeholders, this trend highlights the rising need for streamlined and integrated multi-omic workflow and platform. With Element Biosciences’ AVITI24 system and the only system that can deliver multi-omics data in a single instrument, researchers can fully utilize its capabilities to:
• Generate multi-omics encompassing morphology, protein, RNA targets, and all their respective spatial localization map
• Generate multi-omics data from a single assay
• Carry out multi-omics studies without the need for multiple instrument and ancillary component such as a cell sorter, single cell isolation platform, cell visualization platform, respective consumables for each platform and storage considerations, etc.
Conclusion
ChIP-seq is not a legacy technology; it is an evolving pillar of functional genomics. The market landscape has shifted from instruments that simply generate sequence data to be integrated into a multitude of other systems to solve biological problems.
• Strategic investment in platforms is key in enabling multi-omics projects such as single-cell, spatial, and epigenomics, where the most impactful discoveries in complex disease are now being made.
• The value proposition you should evaluate is not just reads or cost per gigabase, but the end-to-end workflow, application support, and research utility power to turn integrated omics data into translational, actionable biological and therapeutic insights.
The future of gene regulation research is multi-modal, single-cell resolved, and spatially contextual.
The AVITI24™ from Element Biosciences is a state-of-the-art sequencing platform powered by Element’s Avidite Base Chemistry™ (ABC™), which delivers high sequencing accuracy, exceptional data quality, flexible throughput and library prep compatibility, all at an affordable cost.

The AVITI24™ platform, paired with Teton™ CytoProfiling, enables researchers to perform single-cell, spatial multi-omic analysis by co-detecting RNA expression, protein markers, and cell morphology directly from the same sample in a single run, all with subcellular resolution and next-day results.
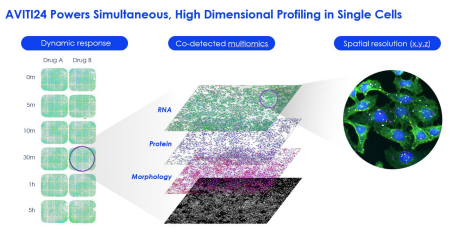
Learn more about AVITI24 and Teton Assays for multi-omics project.


